eplot
Reporting & VisualizationForest plots and coefficient plots from estimates or data
Version 1.1.0 | 2026-04-19
eplot creates forest plots and coefficient plots from three sources — variables in memory, active or stored estimation results, and preassembled matrices — under one command with one set of options.
Quick Start
Run a regression and plot the coefficients in two lines:
sysuse auto, clear
regress price mpg weight foreign
eplot ., drop(_cons) cicap
That's it. eplot . reads the active estimation results and draws a coefficient plot with capped confidence intervals.
Requirements
- Stata 16 or later
- No external package dependencies
Installation
capture ado uninstall eplot
net install eplot, from("https://raw.githubusercontent.com/tpcopeland/Stata-Tools/main/eplot") replace
Commands
| Command | Description |
|---|---|
eplot |
Draw effect plots from data in memory, estimation results, or matrices |
How It Works
eplot detects its input mode automatically from the arguments you supply:
| Call | Mode | When to use |
|---|---|---|
eplot es lci uci, ... |
Data | Effect sizes already in variables (meta-analysis, imported results) |
eplot . or eplot m1 m2, ... |
Estimates | Plot coefficients from a regression you just ran, or compare stored models |
eplot, matrix(R) ... |
Matrix | Results assembled in a Stata matrix |
Mode detection gives precedence to data mode when the first three tokens are numeric variables. In ambiguous cases, use eplot . to force estimates mode or matrix() to force matrix mode explicitly.
Feature Highlights
- One command, three input modes — data in memory, estimation results, and matrices share one option vocabulary
- Journal style presets —
style(lancet),style(jama),style(nejm),style(bmj)apply ready-made looks; override any individual element - Multi-model comparison — overlay stored estimates side by side with
modellabels(),offset(), andpalette() - Values annotation — add formatted effect text next to each marker with
values; customize withvformat()andstars - Significance coloring —
sigcolorsdraws significant and non-significant effects in contrasting colors - Grouped and annotated layouts —
groups(),headers(),gap(), andfavors()organize complex plots - Meta-analysis support — prediction intervals (
pi()), heterogeneity statistics (i2(),tau2(),qstat()), weighted boxes, and diamond pooled effects - Eform with auto-labeling —
eformexponentiates coefficients and sets the axis label automatically (Odds Ratio, Hazard Ratio, IRR)
Worked Examples
1. Coefficient plot with custom labels
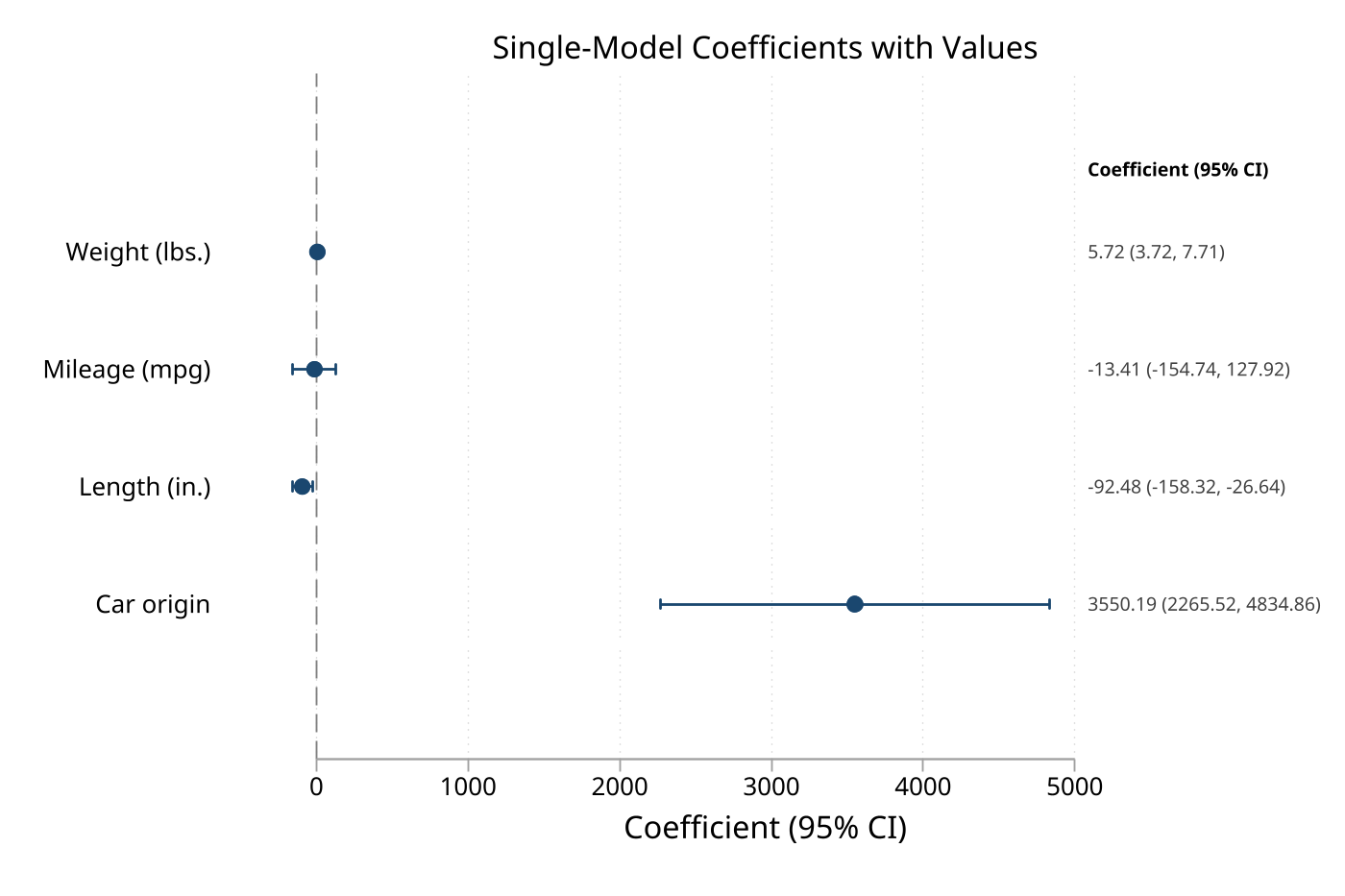
The fastest way to use eplot — run a regression, then plot the results immediately.
sysuse auto, clear
regress price mpg weight foreign
eplot ., drop(_cons) ///
coeflabels(mpg = "Miles per Gallon" ///
weight = "Vehicle Weight" ///
foreign = "Foreign Make") ///
cicap values
2. Compare two stored models
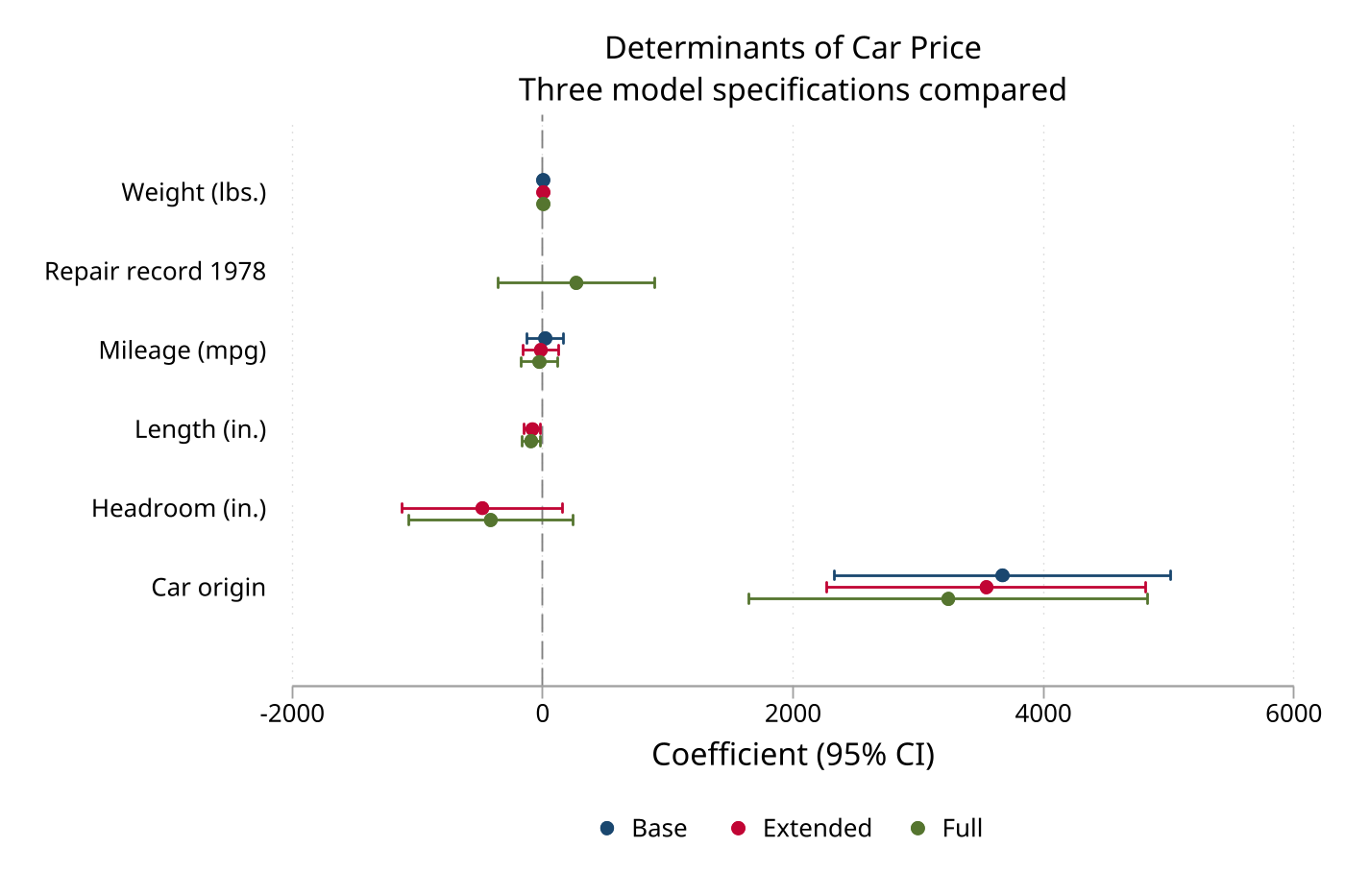
Multi-model estimates mode is useful when you want to compare a base model and an adjusted model side by side.
sysuse auto, clear
quietly regress price mpg weight foreign
estimates store base
quietly regress price mpg weight length foreign headroom
estimates store extended
eplot base extended, drop(_cons) ///
modellabels("Base" "Extended") ///
coeflabels(mpg = "Miles per Gallon" ///
weight = "Vehicle Weight" ///
length = "Body Length" ///
headroom = "Headroom" ///
foreign = "Foreign Make") ///
cicap
3. Forest plot from data in memory
Use data mode when you already have effect sizes and confidence limits in variables — for example, after a meta-analysis or when reading results from another system.
clear
input str20 study es lci uci weight
"Smith 2020" -0.16 -0.36 0.03 15.2
"Jones 2021" -0.33 -0.54 -0.12 18.4
"Brown 2022" -0.09 -0.25 0.06 22.1
"Wilson 2023" -0.39 -0.65 -0.12 12.8
"Overall" -0.24 -0.34 -0.13 .
end
gen byte type = cond(study == "Overall", 5, 1)
eplot es lci uci, labels(study) weights(weight) type(type) ///
values effect("Mean Difference (95% CI)")
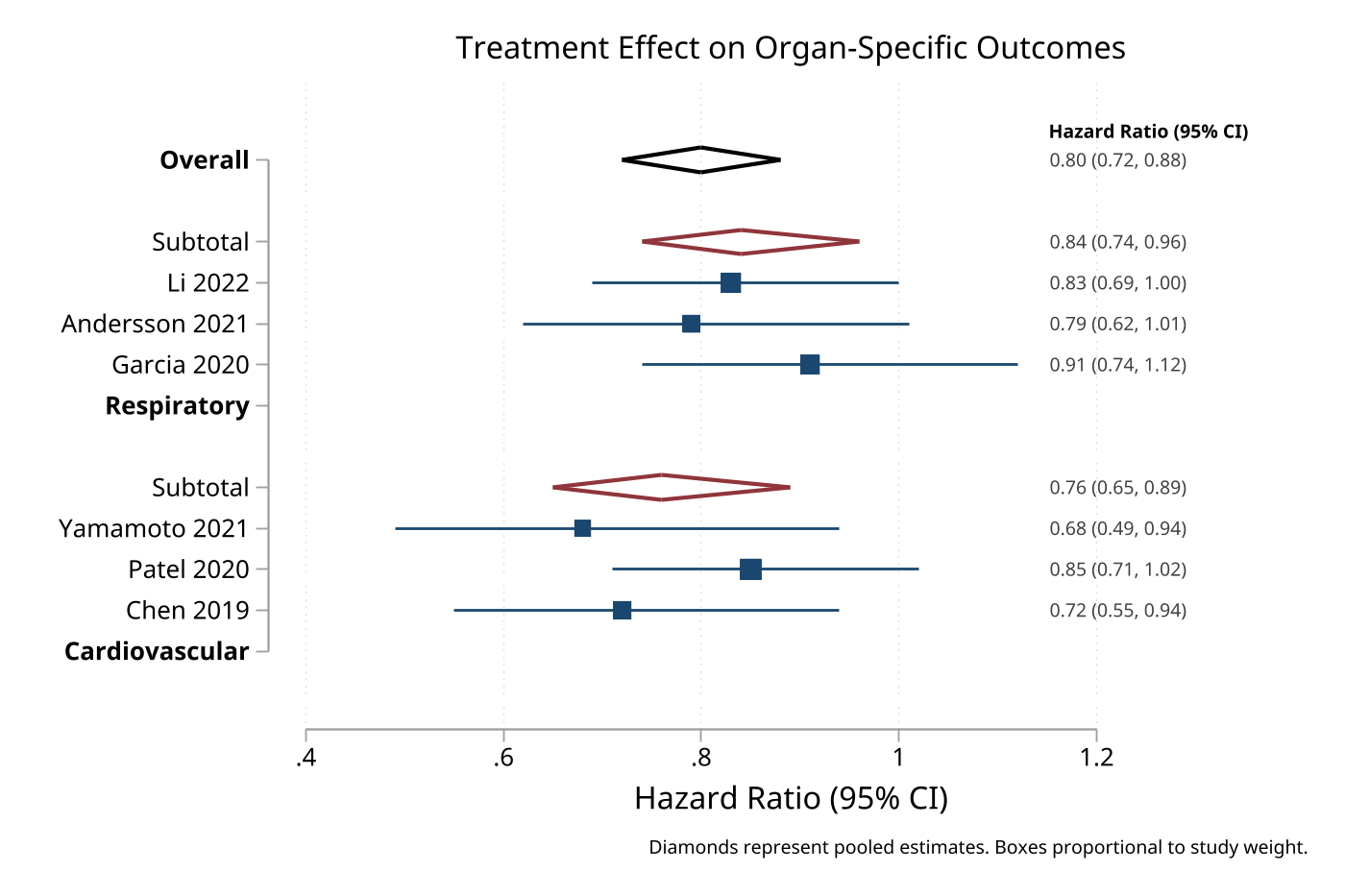
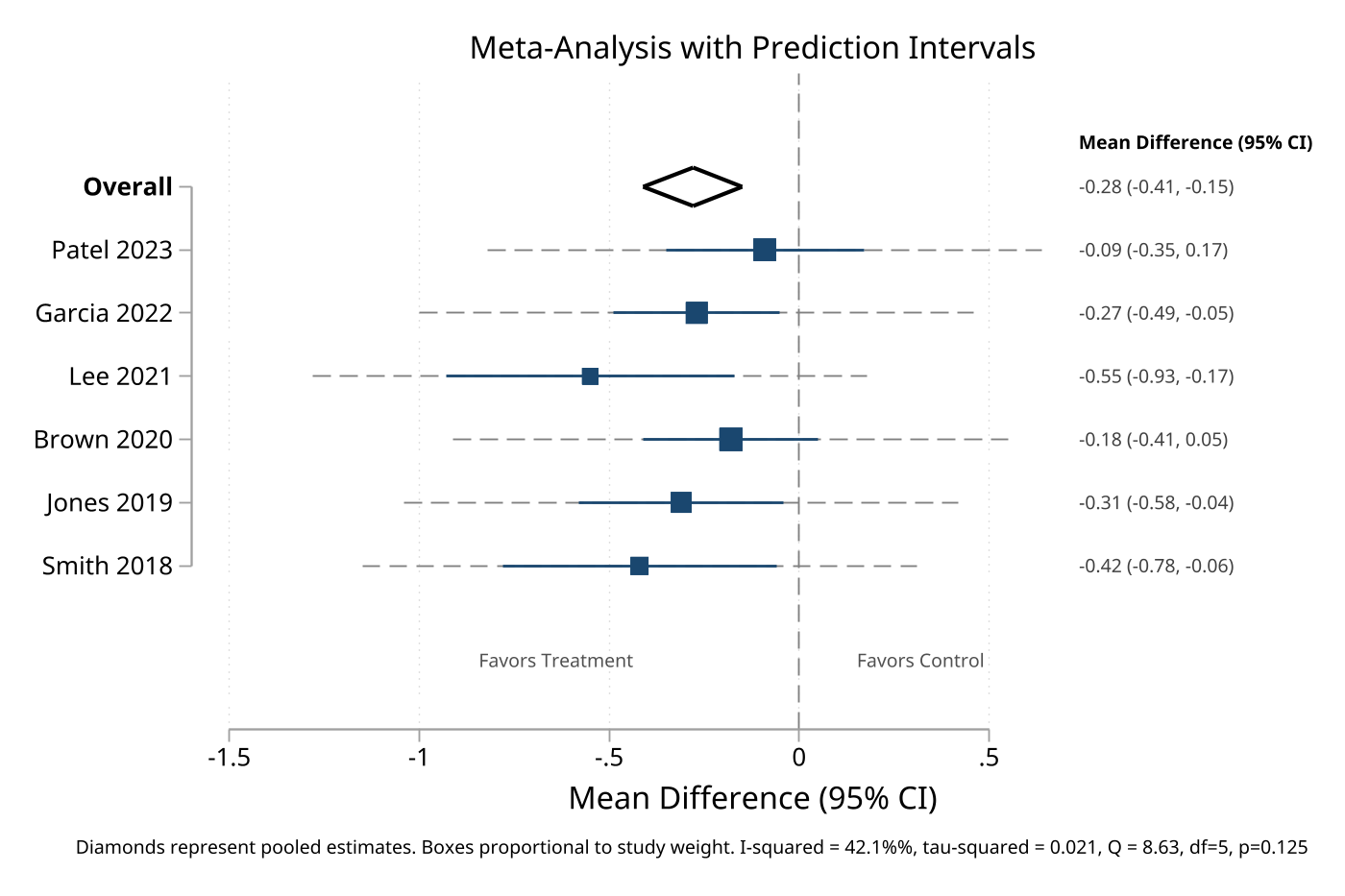
4. Meta-analysis forest plot with heterogeneity
Data mode supports prediction intervals and heterogeneity statistics for publication-ready meta-analysis plots.
clear
input str20 study es lci uci weight byte type
"Smith 2018" -0.42 -0.78 -0.06 12.3 1
"Jones 2019" -0.31 -0.58 -0.04 16.8 1
"Brown 2020" -0.18 -0.41 0.05 21.5 1
"Lee 2021" -0.55 -0.93 -0.17 10.2 1
"Garcia 2022" -0.27 -0.49 -0.05 19.1 1
"Patel 2023" -0.09 -0.35 0.17 20.1 1
"Overall" -0.28 -0.41 -0.15 . 5
end
eplot es lci uci, labels(study) weights(weight) type(type) ///
values vformat(%4.2f) ///
i2("42.1") tau2("0.021") qstat("8.63, df=5, p=0.125") ///
effect("Mean Difference (95% CI)")
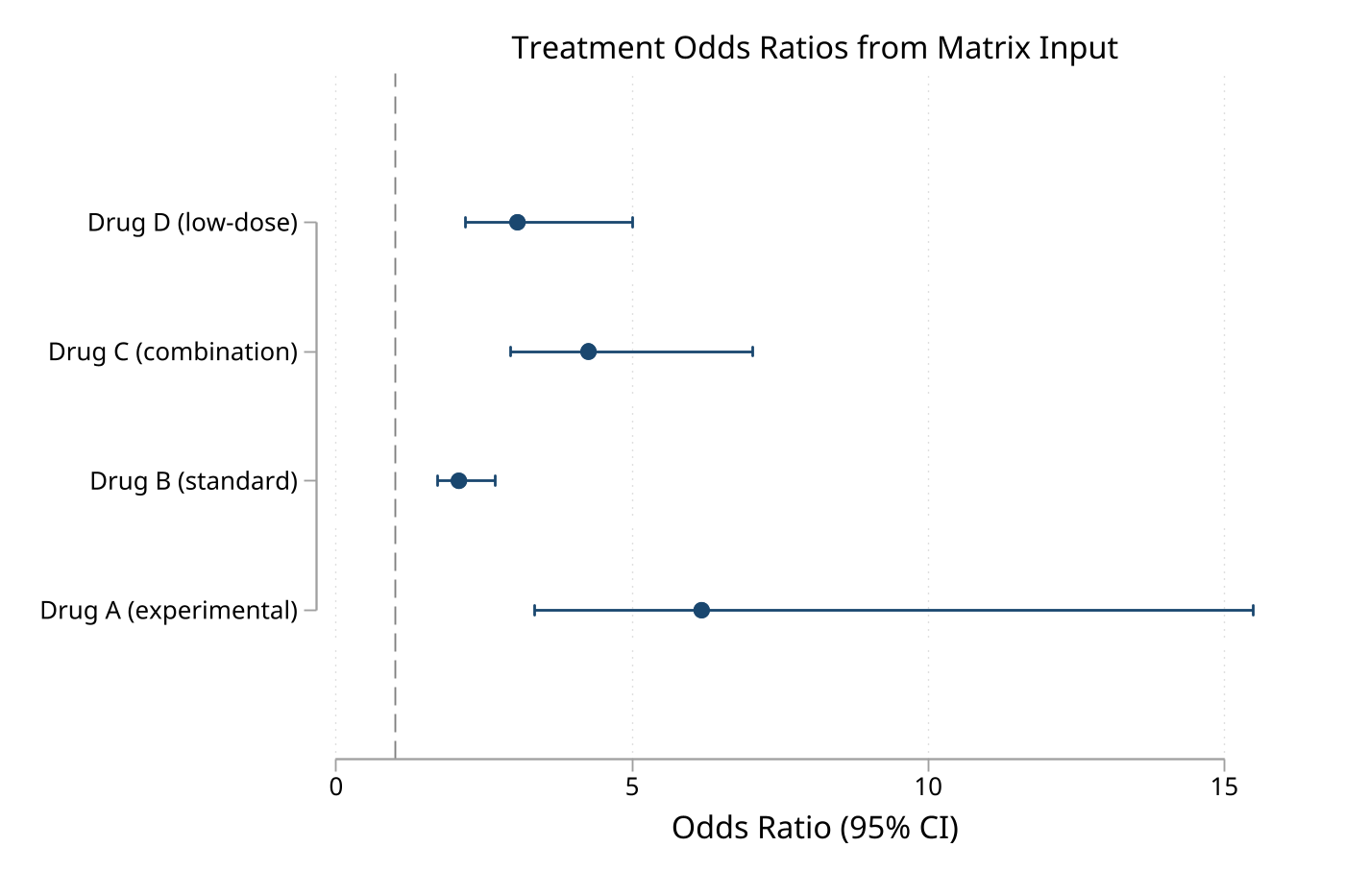
5. Plot from a matrix
Matrix mode is useful when the effect table is already assembled programmatically.
matrix R = (1.5, 1.1, 2.0 \ 0.8, 0.6, 1.2 \ 1.2, 0.9, 1.6)
matrix rownames R = "Treatment_A" "Treatment_B" "Treatment_C"
eplot, matrix(R) eform ///
effect("Odds Ratio") ///
values
6. Logistic regression with odds ratios
After a logistic model, eform exponentiates coefficients to odds ratios and sets the axis label automatically. Factor variables are labeled using Stata's value labels.
sysuse auto, clear
logit foreign mpg weight i.rep78
eplot ., eform cicap values
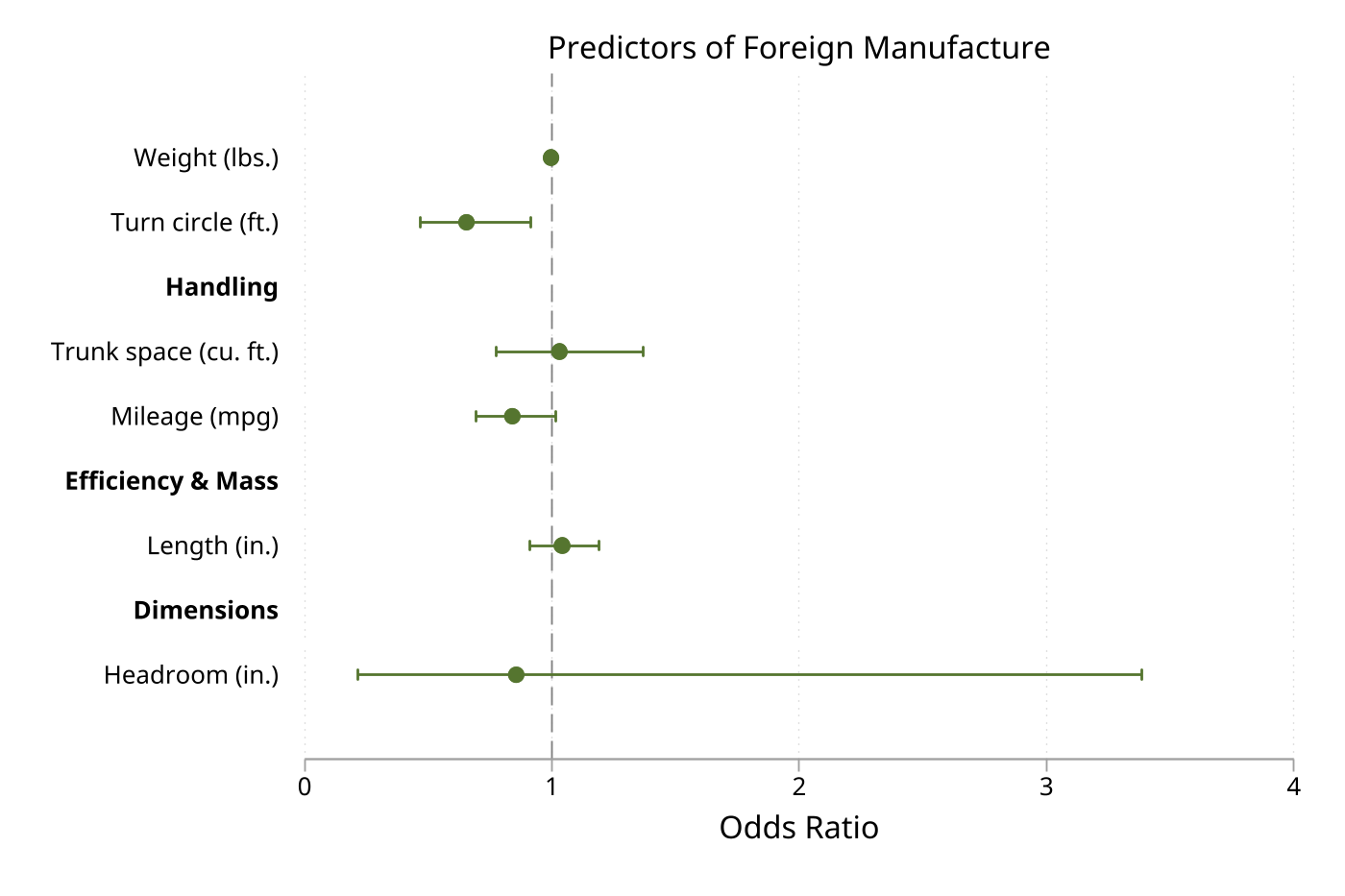
7. Grouped coefficients with significance coloring
Layer options to organize and annotate complex plots. groups() adds section headers, gap() adds space between groups, sigcolors highlights statistical significance, and stars adds asterisks.
sysuse auto, clear
regress price mpg weight length turn foreign rep78
eplot ., noconstant ///
groups(mpg weight length turn = "Vehicle Characteristics" ///
foreign rep78 = "Other Factors") ///
gap(0.5) ///
stars values sigcolors
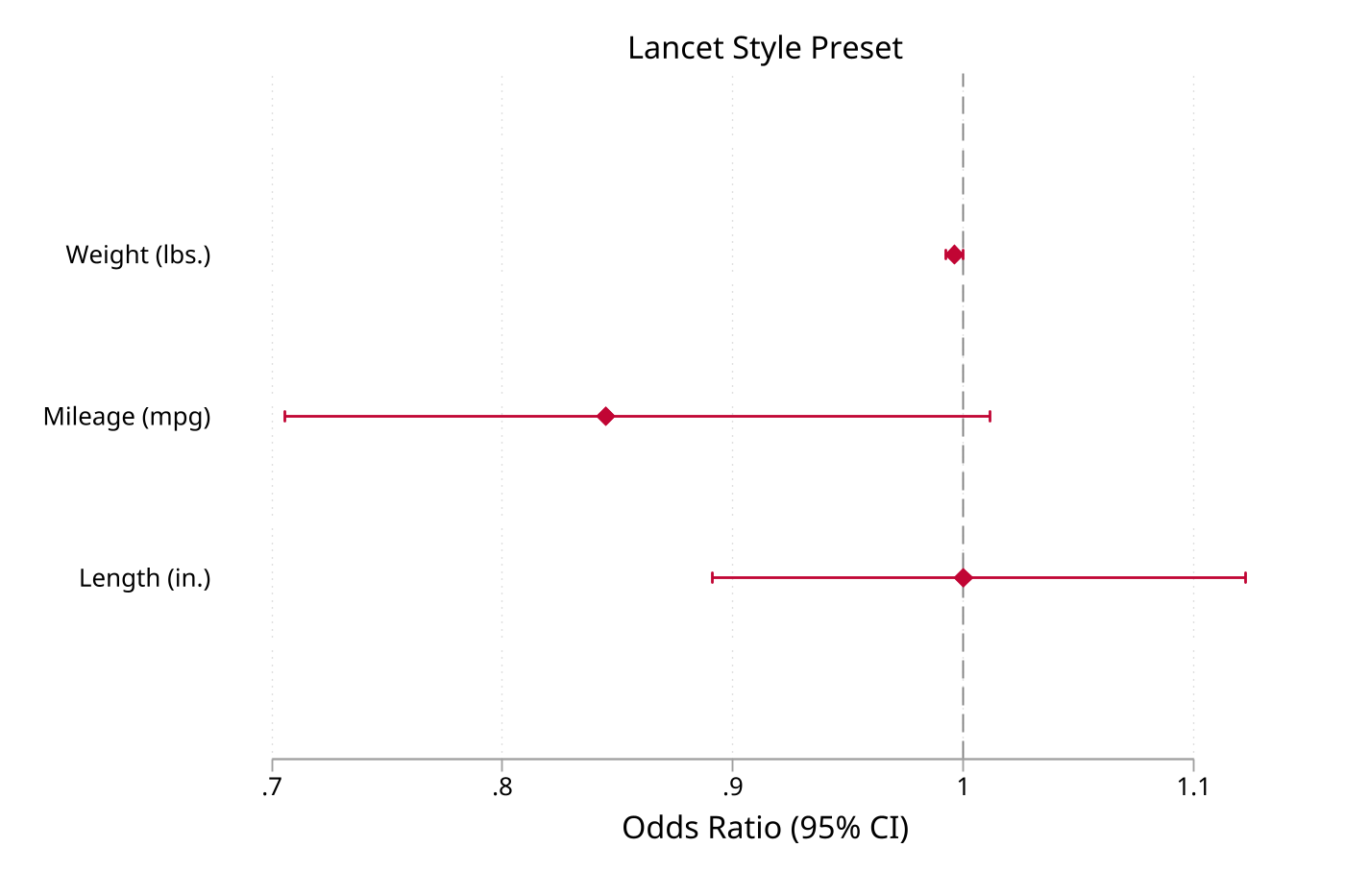
8. Journal style presets
Apply a journal-style preset, then override individual elements as needed.
sysuse auto, clear
regress price mpg weight foreign
eplot ., noconstant style(lancet)
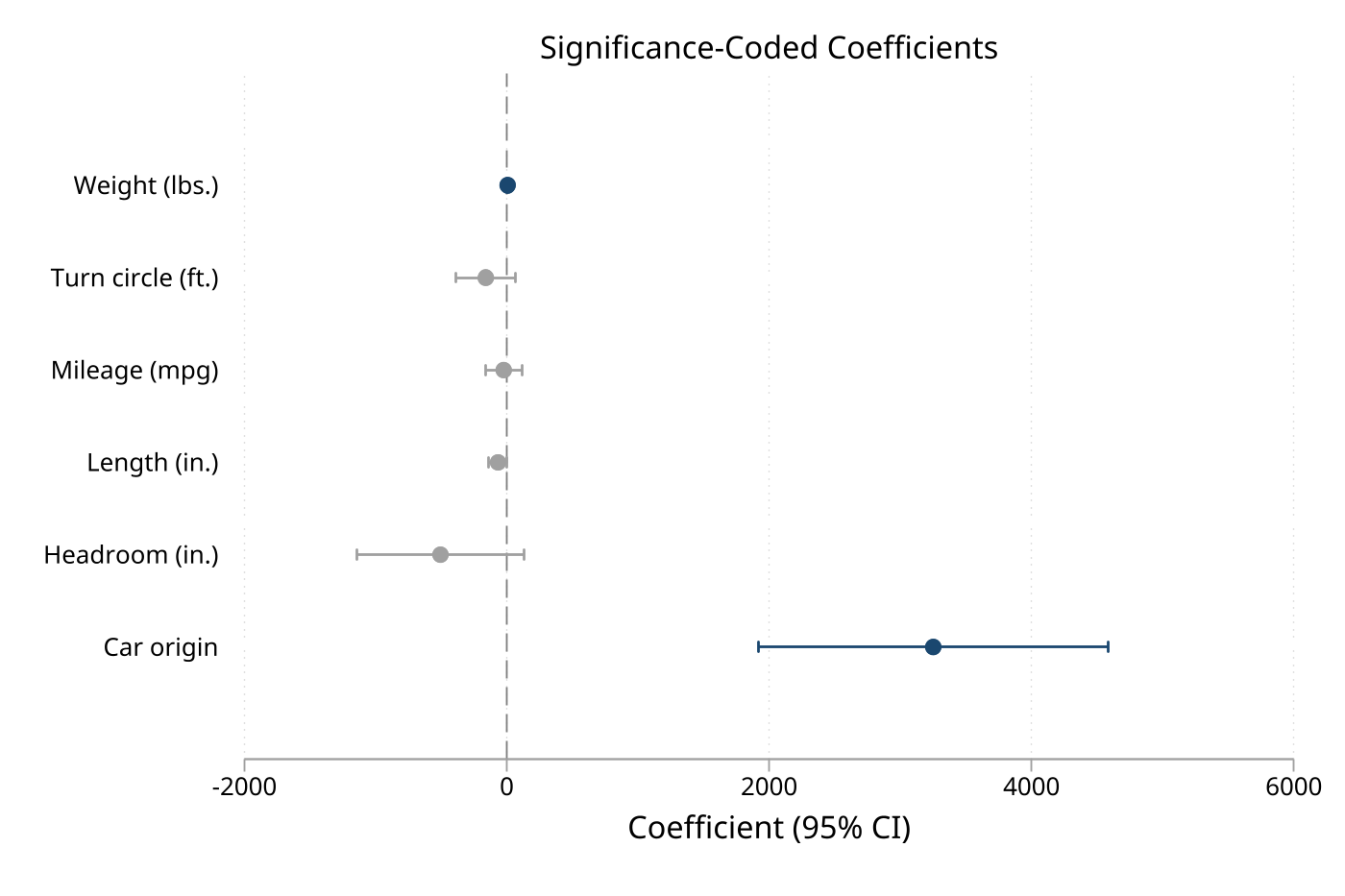
9. Significance coloring with custom colors
eplot ., noconstant sigcolors sigcolor(navy) insigncolor(gs12) cicap
Option Reference
Options are organized by function. Not every option works in every mode — see help eplot for per-option mode availability.
| Category | Options |
|---|---|
| Data specification | labels(), weights(), type() |
| Coefficient selection | keep(), drop(), rename(), noconstant |
| Labeling | coeflabels(), groups(), headers(), gap() |
| Transform | eform, rescale() |
| Reference lines | null(), nonull, xline(), xlabel() |
| Confidence intervals | level(), noci, cicap |
| Display | dp(), effect(), values, vformat(), stars, sigcolors, sigcolor(), insigncolor(), style(), favors() |
| Prediction/heterogeneity | pi(), i2(), tau2(), qstat() |
| Layout | horizontal, vertical, sort, order() |
| Multi-model | modellabels(), offset(), palette(), legendopts() |
| Markers | mcolor(), msymbol(), msize(), boxscale(), nobox, nodiamonds, cicolor(), ciwidth() |
| Graph | title(), subtitle(), note(), name(), saving(), scheme(), plotregion(), graphregion(), aspect(), plus any twoway option |
Also See
help eplot— full documentation with clickable examplescoefplot(Ben Jann, SSC) — comprehensive coefficient plottingmetan(SSC) — meta-analysis with forest plots- Stata 18+:
meta forestplot— official meta-analysis forest plots
Version History
- 1.1.0 (2026-04-19): Added
gap()for grouped spacing, effect-axisxlabel()passthrough, automatic value-annotation margin sizing, and clearer mode-detection documentation - 1.0.0 (2026-04-12): Initial release with data, estimates, and matrix modes; multi-model comparison; journal style presets; significance coloring; meta-analysis features
Author
Timothy P Copeland, Karolinska Institutet