msm
AnalysisMarginal structural models via IPTW for time-varying treatments
Version 1.0.0 | 2026-04-26
msm is a Stata suite for inverse-probability-weighted marginal structural models in person-period data. It is designed for longitudinal settings with time-varying treatments and confounders, where standard regression adjustment can be biased by treatment-confounder feedback.
The package covers the full workflow for conventional static-regime MSM analyses: study protocol specification, variable mapping, validation, stabilized weighting, diagnostics, outcome modeling, counterfactual prediction, plotting, reporting, Excel export, and sensitivity analysis.
When to use this package
Use msm when your data have all of these features:
- Longitudinal panel structure — repeated observations per individual over time.
- Time-varying treatment — treatment status can change between periods.
- Time-varying confounders affected by past treatment — the classic "treatment-confounder feedback" problem. A confounder like biomarker level may be affected by prior treatment and also predict future treatment. Standard regression adjustment cannot handle this without bias; IPTW solves it by reweighting.
- Binary treatment and outcome indicators (0/1). Linear and Cox models are also supported for estimation, but the full prediction workflow requires a binary outcome with a pooled logistic model.
If your treatment is assigned at a single point in time (not time-varying), consider Stata's built-in teffects ipw instead.
Requirements
- Stata 16 or later
Installation
capture ado uninstall msm
net install msm, from("https://raw.githubusercontent.com/tpcopeland/Stata-Tools/main/msm") replace
The release installs msm_example.dta, which the examples below access with findfile.
Commands
Setup and validation
| Command | Description |
|---|---|
msm |
Package overview, workflow guide, and pipeline state check via msm, status |
msm_protocol |
Record the target trial, causal contrast, weighting plan, and analysis plan (7 components) |
msm_prepare |
Map identifier, period, treatment, outcome, censoring, and covariate variables |
msm_validate |
Run 10 data-quality checks for person-period data |
Estimation
| Command | Description |
|---|---|
msm_weight |
Estimate stabilized IPTW and optional censoring weights (IPCW) |
msm_fit |
Fit weighted pooled logistic, linear, or Cox outcome models |
msm_predict |
Generate counterfactual predictions under always-treated and never-treated strategies |
Diagnostics and output
| Command | Description |
|---|---|
msm_diagnose |
Summarize weight distribution and assess covariate balance (SMD before/after) |
msm_plot |
Draw weight density, Love plot, survival curves, trajectory, and positivity plots |
msm_report |
Produce a compact publication-style results table (console, CSV, or Excel) |
msm_table |
Export multi-sheet Excel workbook with all pipeline results |
msm_sensitivity |
Compute E-values and confounding-bound sensitivity summaries |
How It Works
msm is organized as a pipeline. Each step stores its results in the dataset as characteristics, matrices, or variables, and downstream commands read those stored artifacts automatically. This means you only specify your variable mapping once (in msm_prepare) and do not need to repeat it at every step.
The pipeline at a glance
msm_protocol → msm_prepare → msm_validate → msm_weight
↓ ↓
(document) msm_diagnose
↓
msm_fit
↓
msm_predict
↓
msm_plot / msm_report / msm_table / msm_sensitivity
Run msm, status at any point to see the current pipeline stage, what variables are mapped, what artifacts are saved, and what the recommended next step is.
What each step does
msm_protocol— documents the causal question and analysis plan using 7 components adapted from the target trial emulation framework of Hernan et al. (2020). This is purely for documentation; it does not affect computation.msm_prepare— maps your dataset's variable names to roles (ID, period, treatment, outcome, censoring, covariates) and stores the mapping in dataset characteristics. Validates the data structure (person-period format, binary variables, constant baseline covariates). This is the entry point for the analysis.msm_validate— runs 10 data quality checks: person-period format, period gaps, terminal outcomes, treatment variation, missing data, sufficient period sizes, covariate completeness, treatment history patterns, censoring patterns, and positivity by period. Usestrictto treat all warnings as hard errors.msm_weight— fits logistic models for the probability of treatment at each period, then combines the period-specific ratios into cumulative stabilized IP weights. Optionally adds censoring weights (IPCW). Truncation at specified percentiles is available to limit the influence of extreme weights.msm_diagnose— reports the weight distribution (mean, SD, percentiles, effective sample size) and computes standardized mean differences (SMD) for each covariate before and after weighting. A good analysis should see SMDs below 0.1 after weighting.msm_fit— fits the weighted outcome model. The treatment coefficient from this model is the MSM causal estimate. Standard errors are robust/sandwich, clustered at the individual level.msm_predict— generates standardized counterfactual predictions: "What would the outcome be if everyone were always treated? Never treated?" Uses Monte Carlo simulation from the coefficient distribution for confidence intervals. Risk differences between strategies are available.msm_plot,msm_report,msm_table,msm_sensitivity— visualization, reporting, and sensitivity analysis.msm_tableproduces a multi-sheet Excel workbook;msm_reportproduces a single compact summary;msm_sensitivitycomputes E-values for unmeasured confounding.
Choosing an Outcome Model
msm_fit model |
When to use it | Follow-on implications |
|---|---|---|
model(logistic) |
Binary outcomes when you also want standardized counterfactual predictions | Required for msm_predict; use msm, status to confirm prediction is available |
model(linear) |
Continuous outcomes where the weighted mean difference is the target | msm_predict is not available; use msm, status to check the current stage before reporting/export |
model(cox) |
Time-to-event analyses where a weighted hazard ratio is the main estimand | msm_predict is not available; use stcox postestimation and msm, status for pipeline state |
Data Requirements
- Data must be in person-period format, with one row per individual-period.
id()andperiod()must uniquely identify observations.period()must be integer-valued.- All individuals must share a common baseline period before weighting.
treatment()andoutcome()must be binary 0/1 variables.censor()is optional but must also be binary 0/1 when used.- Variables in
baseline_covariates()must be time-fixed (constant within person). msm_weightcurrently rejects delayed entry.msm_predictrequires a priormsm_fit, model(logistic)run.
Interpreting Key Diagnostics
Weight mean
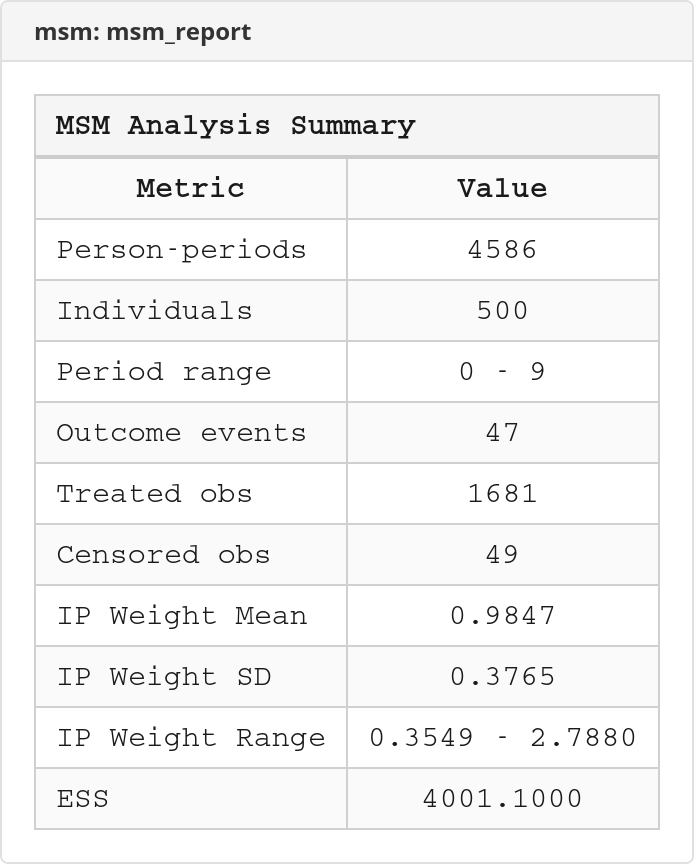
Stabilized IP weights should have a mean close to 1.0. If the mean deviates substantially (e.g., 0.7 or 1.4), the treatment or numerator model may be misspecified. Check your covariate specification.
Effective sample size (ESS)
ESS = (sum of weights)² / (sum of squared weights). It measures how much statistical information the weighted sample retains compared to the original sample. If ESS drops below 50% of N, consider simplifying the weight model or applying stronger truncation.
Standardized mean differences (SMD)
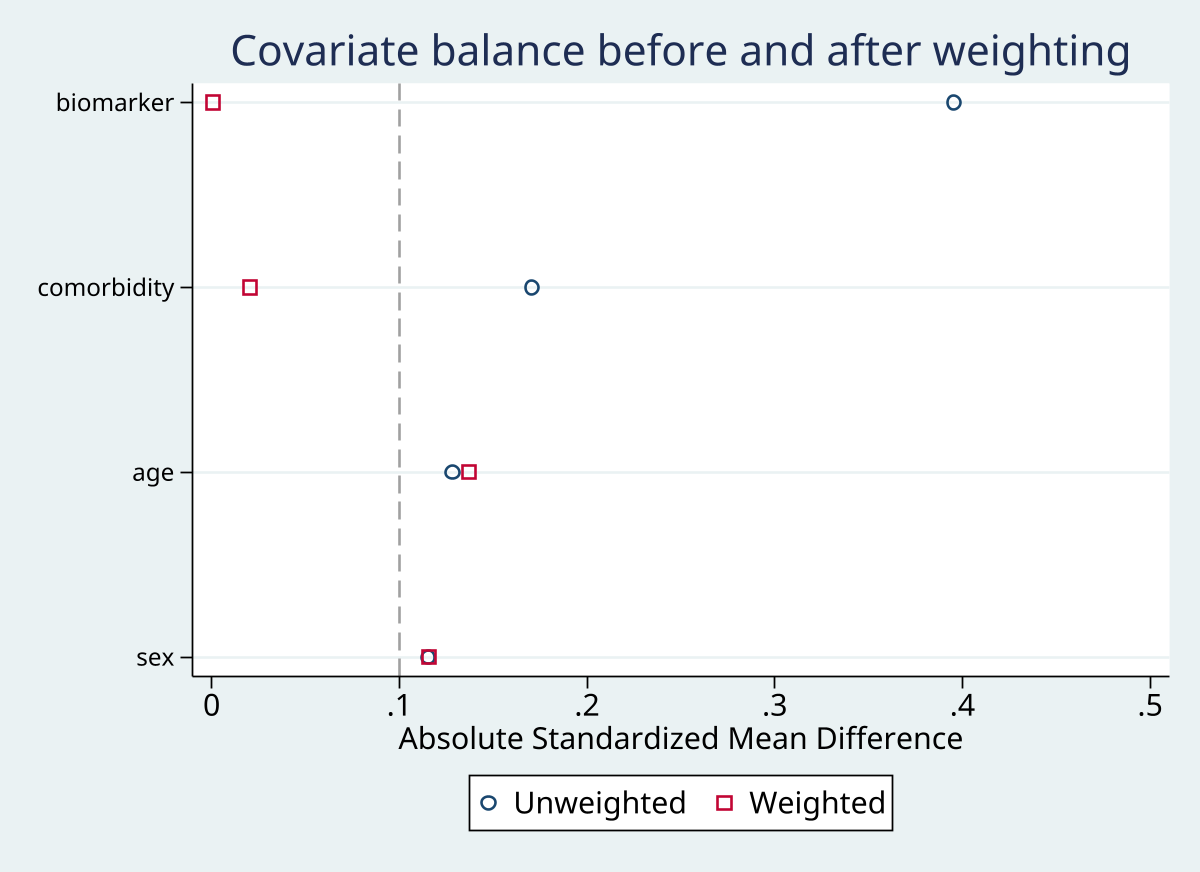
An absolute SMD below 0.1 after weighting is the standard threshold for acceptable covariate balance. SMDs above 0.1 suggest residual confounding for that covariate. If weighting makes balance worse for a variable, investigate the weight model specification.
E-value
The E-value is the minimum strength of association (risk ratio scale) that an unmeasured confounder would need with both treatment and outcome to fully explain away the observed effect. An E-value of 1 means the confidence interval already includes the null. E-values above 2-3 indicate the result is moderately to strongly robust to unmeasured confounding.
Current Scope and Limits
msmtargets static binary treatment strategies. Prediction is implemented for always-treated, never-treated, or both; dynamic and stochastic regimes are not supported.msm_predictrequires a priormsm_fit, model(logistic)run. Linear and Cox fits can be estimated, diagnosed, and reported, but they do not feed intomsm_predict.- If you plan to predict, keep
outcome_cov()limited to covariates that are time-fixed within individual. msm_weightassumes a shared baseline period. Late entry/left truncation is not supported.- By default,
msm_predictonly allowstimes()within the observed follow-up range. Useextrapolateonly when you deliberately want out-of-range predictions.
Demo
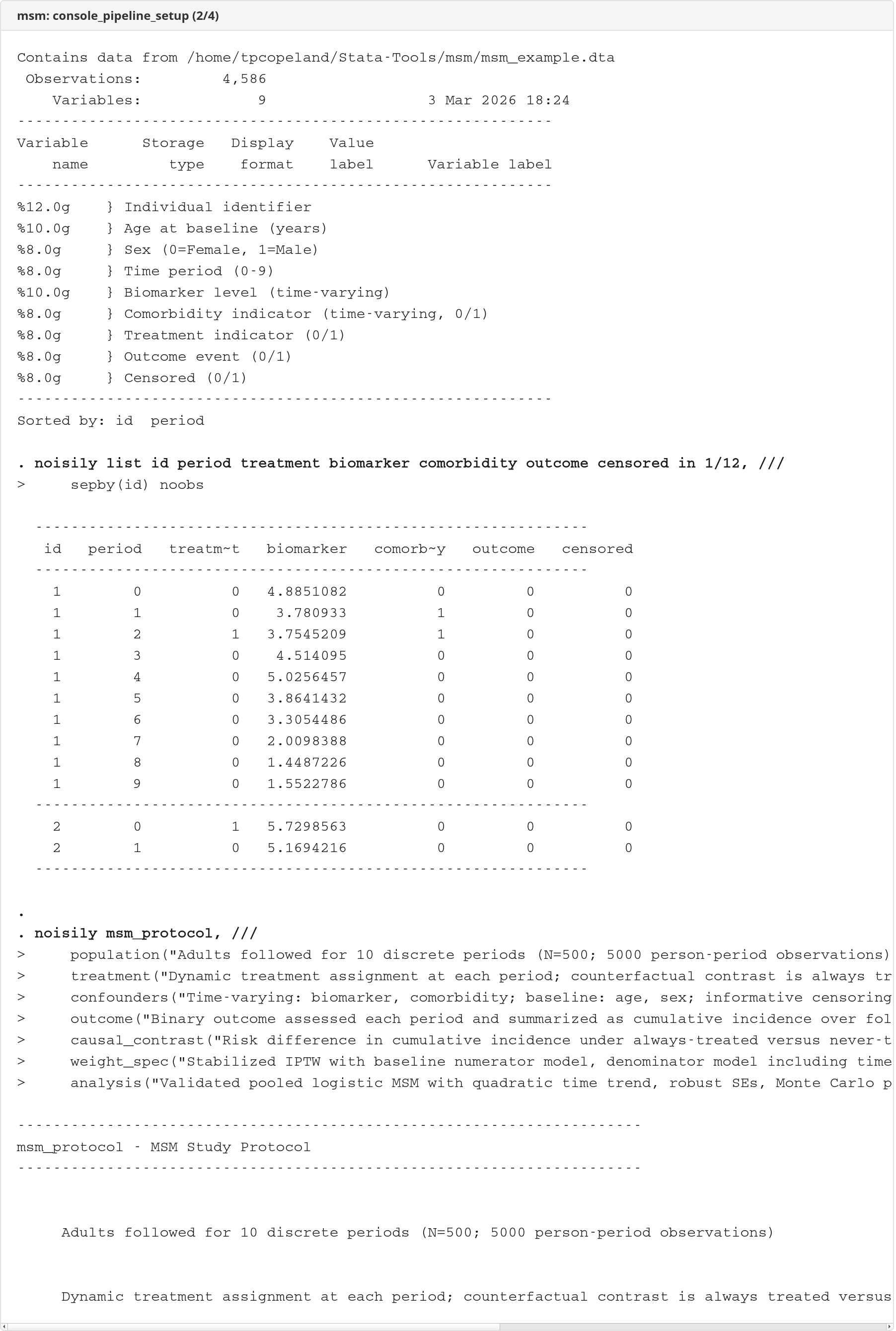
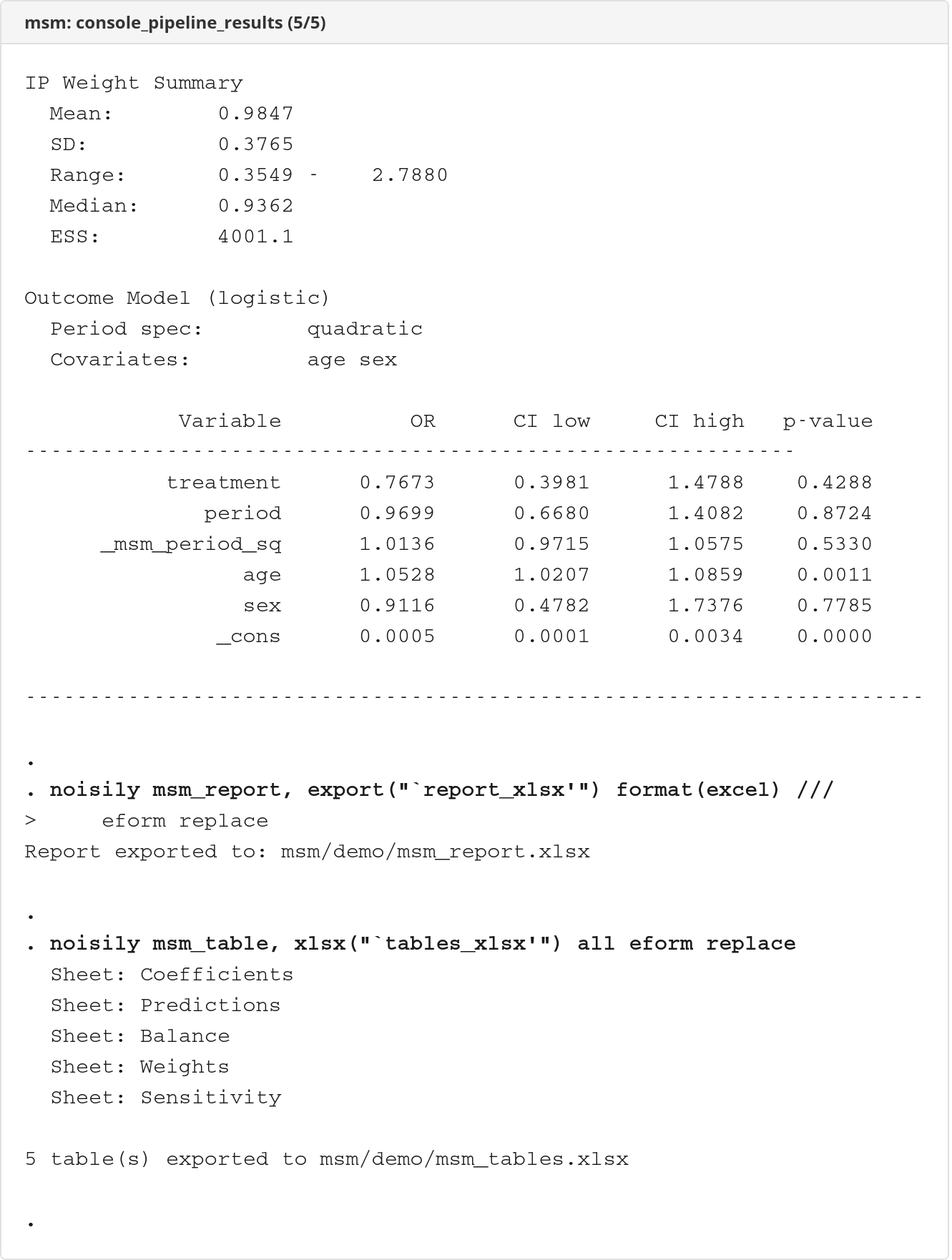
The demo runs the full pipeline on the bundled msm_example.dta dataset. Console output is rendered as self-contained HTML documents using logdoc.
Console output
Graphs
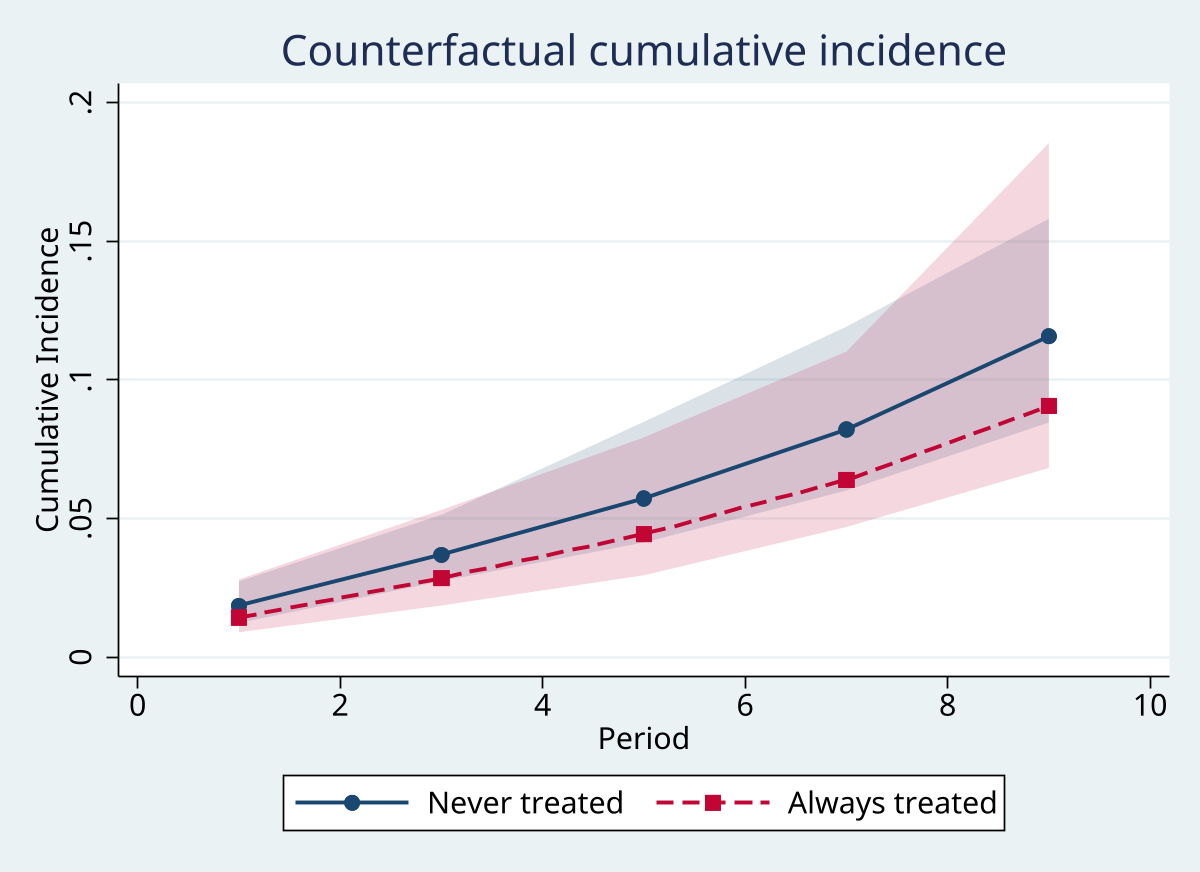
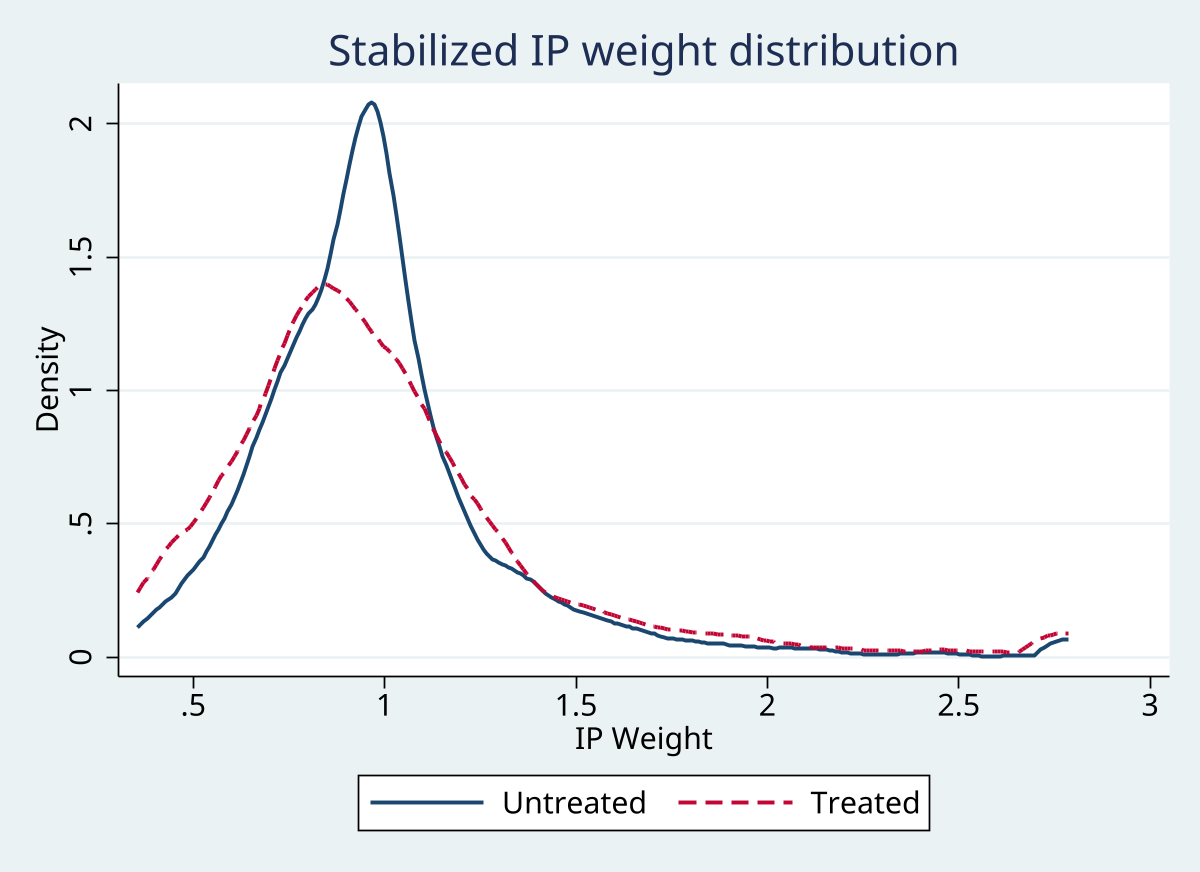
Excel exports
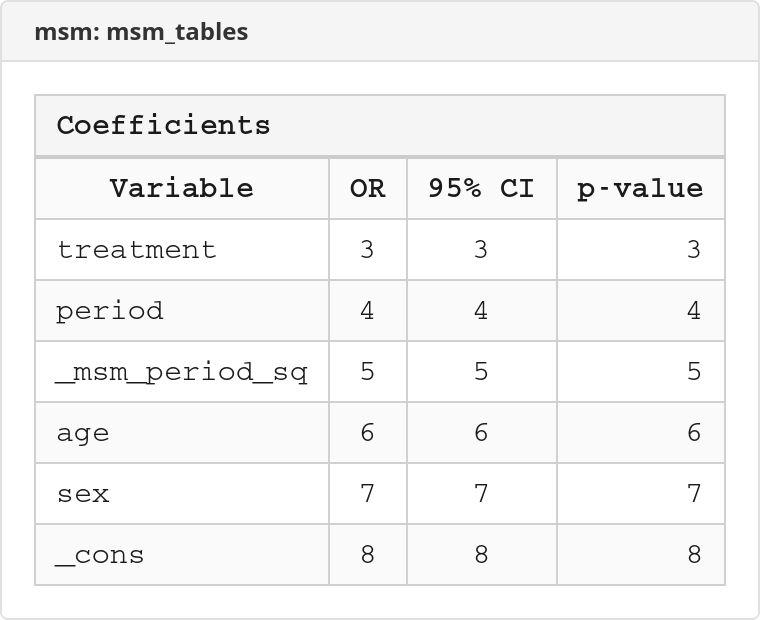
Excel workbook screenshots (click to expand)
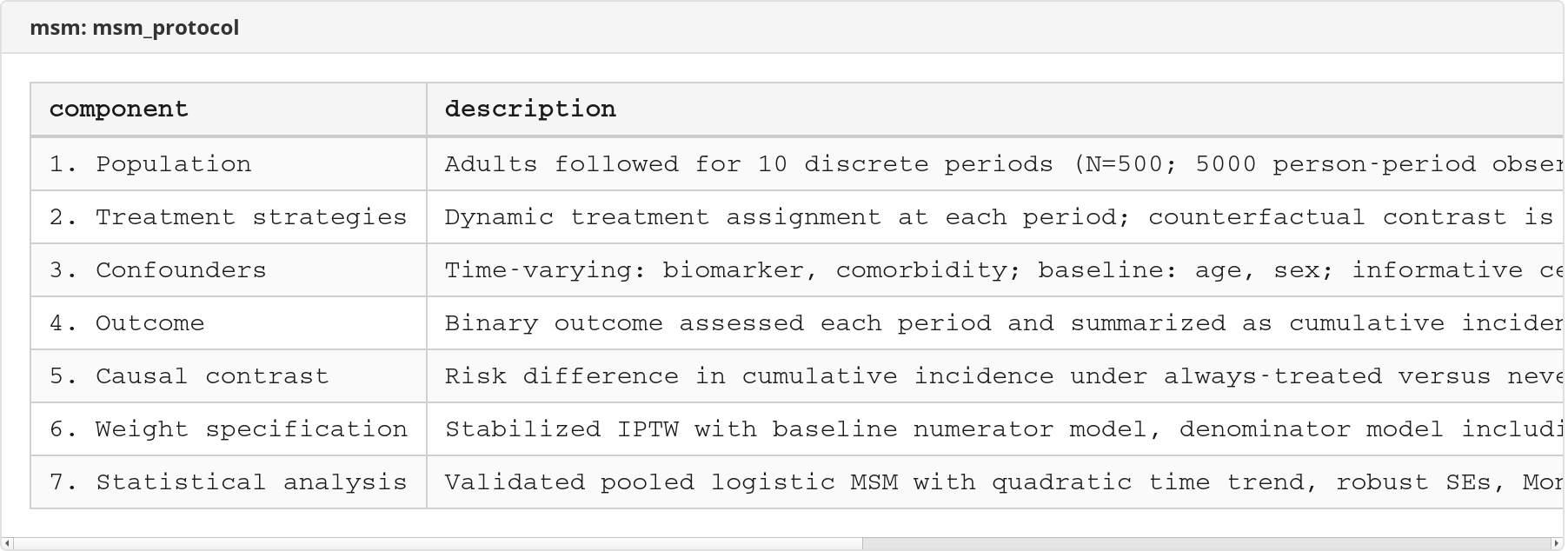

Worked Examples
1. Full pipeline with the bundled example dataset
This example mirrors the package's intended end-to-end workflow. It stays within the supported scope: static always-treat versus never-treat prediction from a pooled logistic MSM.
findfile msm_example.dta
use "`r(fn)'", clear
* Step 0: Document the study protocol
msm_protocol, ///
population("Adults aged 18-65 with condition X") ///
treatment("Always treat vs. never treat") ///
confounders("Biomarker (time-varying), comorbidity (time-varying), age, sex") ///
outcome("Binary clinical endpoint") ///
causal_contrast("ATE: always treat vs. never treat") ///
weight_spec("Stabilized IPTW, truncated at 1st/99th percentile") ///
analysis("Pooled logistic regression, robust SE clustered by ID")
* Step 1: Map variables
msm_prepare, id(id) period(period) treatment(treatment) ///
outcome(outcome) covariates(biomarker comorbidity) ///
baseline_covariates(age sex)
* Step 2: Validate data quality
msm_validate, strict verbose
* Step 3: Calculate stabilized IP weights
msm_weight, treat_d_cov(biomarker comorbidity age sex) ///
treat_n_cov(age sex) truncate(1 99) nolog
* Step 4: Diagnose weights and balance
msm_diagnose, balance_covariates(biomarker comorbidity age sex) ///
by_period threshold(0.1)
* Step 5: Fit the weighted outcome model
msm_fit, model(logistic) outcome_cov(age sex) nolog
* Check pipeline state
msm, status
* Step 6: Counterfactual predictions
msm_predict, times(1 3 5 7 9) type(cum_inc) difference ///
samples(200) seed(12345)
* Step 7: Sensitivity analysis
msm_sensitivity, evalue
* Step 8: Reporting and visualization
msm_plot, type(survival) times(1 3 5 7 9) seed(12345)
msm_report, eform
msm_table, xlsx(msm_results.xlsx) all eform replace
2. Minimal estimation-and-prediction workflow
If you want the core causal estimates first, this shorter sequence gets you from prepared data to standardized counterfactual predictions quickly.
findfile msm_example.dta
use "`r(fn)'", clear
msm_prepare, id(id) period(period) treatment(treatment) ///
outcome(outcome) covariates(biomarker comorbidity) ///
baseline_covariates(age sex)
msm_validate
msm_weight, treat_d_cov(biomarker comorbidity age sex) ///
treat_n_cov(age sex) nolog
msm_fit, model(logistic) outcome_cov(age sex) nolog
msm, status
msm_predict, times(3 5 7 9) difference seed(12345)
3. Estimation-only workflow (Cox model)
When the target estimand is a weighted hazard ratio and prediction is not needed:
findfile msm_example.dta
use "`r(fn)'", clear
msm_prepare, id(id) period(period) treatment(treatment) ///
outcome(outcome) covariates(biomarker comorbidity) ///
baseline_covariates(age sex)
msm_weight, treat_d_cov(biomarker comorbidity age sex) ///
treat_n_cov(age sex) nolog
msm_fit, model(cox) outcome_cov(age sex) nolog
msm_report, eform
Output Notes
msm_weightcreates_msm_weightand returns weight summaries such asr(mean_weight),r(ess), andr(n_truncated).msm_diagnosereturns a balance matrix inr(balance)whenbalance_covariates()is specified.msm_fitstores the weighted model ine()and records the fitted MSM effect matrix ine(effects).msm_predictreturns the prediction matrix inr(predictions), risk differences inr(rd_#)whendifferenceis requested, and the seed/state used for the Monte Carlo draws inr(seed)plusr(seed_state).msm_tableexports formatted Excel workbooks;msm_reportproduces compact summaries to console, CSV, or Excel.
References
- Robins JM, Hernan MA, Brumback B. Marginal structural models and causal inference in epidemiology. Epidemiology. 2000;11(5):550-560.
- Hernan MA, Brumback B, Robins JM. Marginal structural models to estimate the causal effect of zidovudine on the survival of HIV-positive men. Epidemiology. 2000;11(5):561-570.
- Cole SR, Hernan MA. Constructing inverse probability weights for marginal structural models. American Journal of Epidemiology. 2008;168(6):656-664.
- VanderWeele TJ, Ding P. Sensitivity analysis in observational research: introducing the E-value. Annals of Internal Medicine. 2017;167(4):268-274.
- Hernan MA, Robins JM. Causal Inference: What If. Boca Raton: Chapman & Hall/CRC, 2020.
Version History
- 1.0.0 (2026-04-26): Initial Stata-Tools release of the full MSM workflow suite
Author
Timothy P Copeland, Karolinska Institutet
License
MIT