psdash
AnalysisPropensity score diagnostics dashboard (overlap, balance, weights, support)
Version 1.0.0 | 2026-04-29
Unified diagnostics dashboard for propensity score analyses in Stata. After teffects, after logit/probit with manually supplied propensity scores from predict, or in fully manual mode, psdash assesses the four standard PS diagnostic domains through one command family: overlap between treatment groups (psdash overlap), covariate balance before and after weighting (psdash balance), weight distribution and effective sample size (psdash weights), and common-support regions (psdash support). psdash combined runs all four and produces a consolidated dashboard.
This package exists because PS diagnostics in Stata are scattered across tebalance, user-written helpers, and ad-hoc summarize/tabstat calls, with each step requiring the analyst to re-specify the treatment variable, covariate list, PS variable, and weighting scheme. psdash collapses that friction: when called after teffects, it reads treatment, covariates, propensity scores, and the implied weighting scheme (ATE/ATT/ATC) directly from e(). After logit/probit it still pulls treatment and covariates from the estimation context. In fully manual mode, treatment and PS are passed explicitly; covariates() and wvar() are supplied to the subcommands that use those inputs. Auto-generated propensity scores and IPTW weights are created as temporary working variables and are not left behind in the user's dataset.
Balance reporting is deliberately richer than the tebalance summarize default. psdash balance computes raw and weighted standardized mean differences, variance ratios, Kolmogorov-Smirnov statistics, and a Love plot sorted by absolute SMD, with configurable thresholds and Excel export. When a PS is available, it auto-generates IPTW weights for the requested estimand() (default ATE) and displays adjusted columns alongside raw columns, so the user sees immediately how much weighting resolves any imbalance — with nowvar to suppress weighting and wvar() to supply a pre-computed weight variable. Factor and interaction notation (i.var, c.var, ##) is expanded transparently when balance is auto-detected from a fitted logit/probit/teffects model.
The weights subcommand is the complement. psdash weights reports mean, SD, range, percentiles, effective sample size, and extreme-weight counts, with on-the-fly trim(#), truncate(#), and stabilize modifications exposed through generate(name) so the modified weights are kept as a new variable rather than overwriting the original. psdash support assesses common-support regions via manual PS thresholds or the Crump et al. (2009) optimal-trimming rule and can write an in_support indicator for downstream analyses. All subcommands store results in r() and the dashboard output lines use clear status labels plus a "Consider:" action line when follow-up is warranted.
Installation
* Released version from GitHub:
capture ado uninstall psdash
net install psdash, from("https://raw.githubusercontent.com/tpcopeland/Stata-Tools/main/psdash") replace
* Development install from a local checkout:
capture ado uninstall psdash
net install psdash, from("/path/to/psdash") replace
How It Works
psdash is designed to work in four modes:
- After
teffects: treatment, covariates, propensity scores, and the implied weighting scheme are auto-detected frome(). This is the shortest workflow: fitteffects, then runpsdash combinedor one of the individual subcommands. - After
logit/probit: treatment and covariates are read from the estimation context, but you still supply the PS variable created bypredict. - After
mlogit(multi-group): for multi-valued treatments, treatment and covariates are auto-detected frome(). Runpredict ps1 ps2 ps3, prand pass the GPS variables viapsvars(ps1 ps2 ps3). - Manual mode: provide treatment and PS explicitly, then pass
covariates()to balance/combined andwvar()to balance/weights/combined when you want to override auto-detection.
When a PS variable is available, psdash balance auto-generates IPTW weights for the requested estimand() unless you suppress that with nowvar or provide wvar() yourself.
Auto-generated propensity scores and weights are temporary working variables and are not left behind in the user's dataset.
Worked Examples
All examples below use Stata's built-in sysuse or webuse datasets, so they can be copied directly after installation. The sysuse auto examples use foreign as a convenient binary treatment indicator. The webuse cattaneo2 examples show a more realistic treatment-effects workflow. They are intended to illustrate syntax and diagnostics, not to endorse a final causal specification.
1. Manual propensity-score workflow with sysuse auto
Estimate the propensity score with logit, save fitted probabilities in ps, then run each diagnostic explicitly. Because this is a manual workflow, balance is told which covariates to assess.
sysuse auto, clear
logit foreign mpg weight length
predict double ps, pr
psdash overlap foreign ps
psdash balance foreign ps, covariates(mpg weight length) loveplot
psdash weights foreign ps
psdash support foreign ps, crump generate(in_support)
2. Fully automatic workflow after teffects with webuse cattaneo2
Here teffects ipw estimates the propensity score internally. After that, psdash reads the treatment (mbsmoke), the estimated PS, the covariates, and the implied weighting scheme from e(), so the subcommands can be called without retyping variable names.
webuse cattaneo2, clear
teffects ipw (bweight) (mbsmoke mage prenatal1 mmarried fbaby)
psdash combined
psdash balance
3. Using pre-computed weights with sysuse auto
If weights were created outside psdash, pass them through wvar() so that the weight and balance diagnostics use the same variable.
sysuse auto, clear
logit foreign mpg weight length
predict double ps, pr
gen double ipw = cond(foreign == 1, 1/ps, 1/(1-ps))
psdash weights foreign ps, wvar(ipw) detail graph
psdash balance foreign ps, covariates(mpg weight length) wvar(ipw)
4. ATT workflow after teffects, atet
When teffects is fit with , atet, psdash maps Stata's ATET result to estimand(att) internally. That lets the balance and weight diagnostics use ATT weights automatically.
webuse cattaneo2, clear
teffects ipw (bweight) (mbsmoke mage prenatal1 mmarried fbaby), atet
psdash balance
psdash weights, detail
5. Focused option examples on sysuse auto
These examples show common follow-up diagnostics once ps already exists: add KS statistics to the balance table, create trimmed or stabilized weights, and mark observations inside common support.
sysuse auto, clear
logit foreign mpg weight length
predict double ps, pr
psdash balance foreign ps, covariates(mpg weight length) ks
psdash weights foreign ps, trim(99) generate(ipw_trimmed)
psdash weights foreign ps, stabilize generate(ipw_stab)
psdash support foreign ps, crump generate(in_support)
Subcommands
| Subcommand | Purpose |
|---|---|
overlap |
PS density/histogram by treatment group |
balance |
SMD balance table + Love plot |
weights |
Weight distribution, ESS, extreme weights, trim/stabilize |
support |
Common support assessment, Crump optimal trimming |
combined |
All diagnostics in a combined dashboard |
Key Options
balance
covariates(varlist)— covariates to assess (auto-detected if omitted)wvar(varname)— weight variable (auto-generated from PS if omitted)
Default behavior: When a PS is supplied,
balanceauto-generates IPTW weights for the requestedestimand()(default: ATE) and displays adjusted SMD/VR columns alongside the raw columns. Passnowvarto see raw balance only, orwvar()to supply a pre-computed weight variable.
loveplot— generate Love plotthreshold(#)— SMD threshold (default: 0.1)xlsx(filename)— export to Excel
weights
wvar(varname)— weight variable (auto-generated from PS if omitted)trim(#)— trim at percentile (50–99.9)truncate(#)— cap at fixed valuestabilize— create stabilized weightsgenerate(name)— variable for modified weightsdetail— show percentile distributiongraph— weight distribution histogram
support
crump— Crump et al. (2009) optimal trimmingthreshold(#)— manual PS trimming threshold (0–0.5)generate(name)— create in-support indicator
combined
nooverlap,nobalance,noweights,nosupport— suppress panels
Stored Results
Each subcommand stores results in r(). See help psdash for the full list.
Relationship to Existing Packages
psdash balance incorporates the computation from balancetab and psdash weights from iptw_diag. Both use identical methods. The overlap and support subcommands are new.
Demo
Binary treatment (2 groups)
Synthetic data: 800 observations, confounded treatment assignment (statin use), propensity scores via logit, IPTW weights.
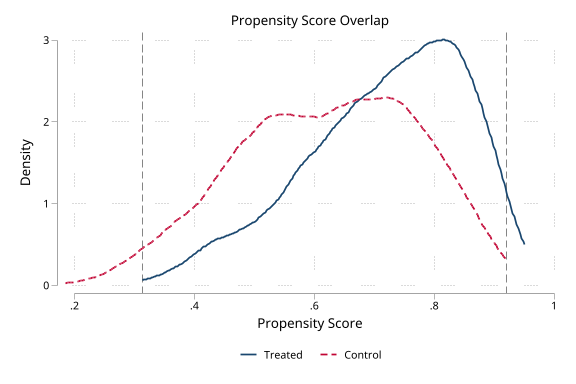
Overlap diagnostics (click to expand)
. noisily psdash overlap statin ps, nograph
----------------------------------------------------------------------
Propensity Score Overlap
----------------------------------------------------------------------
Treatment: statin
PS variable: ps
----------------------------------------------------------------------
----------------------------------------------------------------------
Propensity Score Distribution
----------------------------------------------------------------------
Treated Control
----------------------------------------------------------------------
N 551 249
Mean 0.7191 0.6217
SD 0.1325 0.1486
Min 0.3136 0.1853
Max 0.9505 0.9206
----------------------------------------------------------------------
-------------------------------------------------------
Common Support Region
-------------------------------------------------------
Lower bound: 0.3136
Upper bound: 0.9206
Outside support: 17 ( 2.12%)
Treated outside: 12
Control outside: 5
C-statistic (AUC): 0.6881
-------------------------------------------------------
Overlap: Good ( 2.1% outside support)
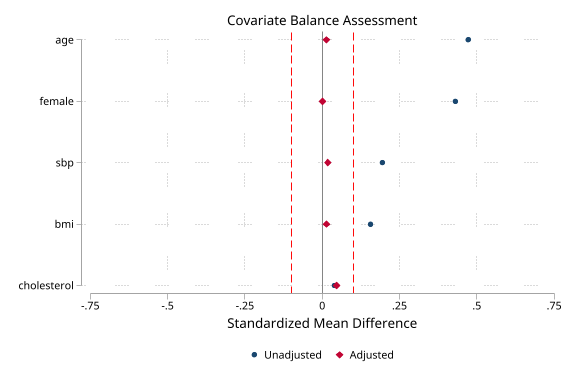
Balance and weight diagnostics (click to expand)
. noisily psdash balance statin ps,
> covariates(age female bmi sbp cholesterol) wvar(ipw)
---------------------------------------------------------------------------
Covariate Balance Assessment
---------------------------------------------------------------------------
Treatment: statin
Estimand: ATE
N (treated): 551
N (control): 249
Weights: ipw
Threshold: 0.100
---------------------------------------------------------------------------
---------------------------------------------------------------------------------------
Covariate | SMD Raw VR Raw SMD Adj VR Adj Status
---------------------------------------------------------------------------------------
age | 0.472 0.99 0.013 1.01 Balanced
female | 0.430 1.03 0.001 1.00 Balanced
bmi | 0.156 1.02 0.014 1.07 Balanced
sbp | 0.194 1.01 0.018 1.05 Balanced
cholesterol | 0.039 1.04 0.047 0.99 Balanced
---------------------------------------------------------------------------------------
Maximum |SMD| (raw): 0.472
Maximum |SMD| (adjusted): 0.047
Maximum VR (raw): 1.04
Maximum VR (adjusted): 1.07
Covariates > SMD threshold: 0 of 5
---------------------------------------------------------------------------------------
Balance: Adequate (max |SMD| = 0.047)
. noisily psdash weights statin ps, wvar(ipw)
----------------------------------------------------------------------
IPTW Weight Diagnostics
----------------------------------------------------------------------
Weight variable: ipw
Treatment: statin
Observations: 800
----------------------------------------------------------------------
----------------------------------------------------------------------
Weight Distribution Summary
----------------------------------------------------------------------
Overall Treated Control
----------------------------------------------------------------------
N 800 551 249
Mean 1.996 1.450 3.206
SD 1.271 0.334 1.682
Min 1.052 1.052 1.227
Max 12.602 3.188 12.602
----------------------------------------------------------------------
----------------------------------------------------------------------
Effective Sample Size (ESS)
----------------------------------------------------------------------
Overall Treated Control
----------------------------------------------------------------------
ESS 569.4 523.2 195.4
ESS % of N 71.2% 95.0% 78.5%
----------------------------------------------------------------------
--------------------------------------------------
Extreme Weight Detection
--------------------------------------------------
Coefficient of Variation: 0.637
Weights > 10: 3 ( 0.38%)
Weights > 20: 0
--------------------------------------------------
Warning: 3 extreme weights detected (>10).
Weights: Acceptable (ESS = 71.2% of N)
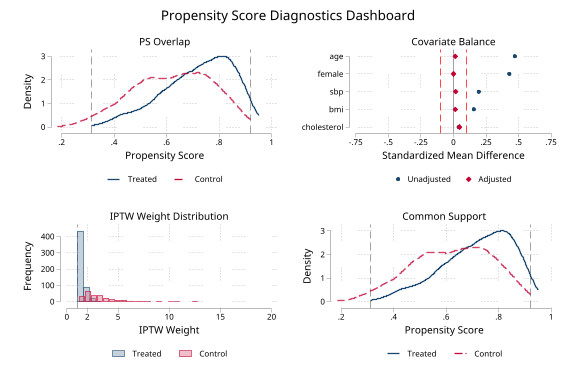
Common support assessment (click to expand)
. noisily psdash support statin ps, crump nograph
----------------------------------------------------------------------
Common Support Assessment
----------------------------------------------------------------------
Treatment: statin
PS variable: ps
Observations: 800
----------------------------------------------------------------------
------------------------------------------------------------
Propensity Score Range
------------------------------------------------------------
Treated Control
------------------------------------------------------------
N 551 249
Min PS 0.3136 0.1853
Max PS 0.9505 0.9206
------------------------------------------------------------
-------------------------------------------------------
Common Support Region
-------------------------------------------------------
Lower bound: 0.3136
Upper bound: 0.9206
Outside support: 17 ( 2.12%)
Treated outside: 12
Control outside: 5
-------------------------------------------------------
-------------------------------------------------------
Crump et al. (2009) Optimal Trimming
-------------------------------------------------------
Optimal alpha: 0.1000
Trim region: [0.100, 0.900]
Observations trimmed: 26 ( 3.25%)
Remaining sample: 774
-------------------------------------------------------
Support: Trimmed ( 3.2% excluded)
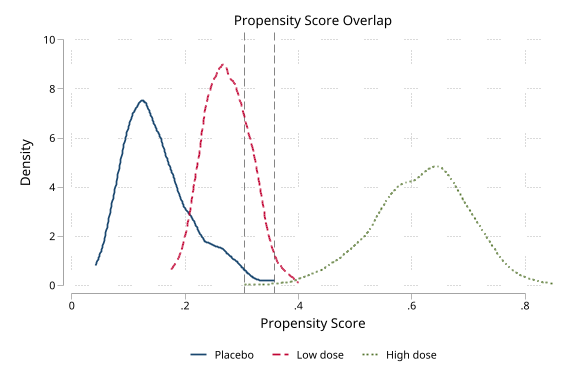
Multi-group treatment (3 arms)
Synthetic data: 1,200 observations, 3-arm treatment (placebo / low dose / high dose) assigned via multinomial logit with confounding by age, BMI, and SBP. Generalized propensity scores via mlogit, generalized IPTW weights.
Multi-group overlap (click to expand)
. noisily psdash overlap arm, psvars(ps0 ps1 ps2) nograph
----------------------------------------------------------------------
Propensity Score Overlap
----------------------------------------------------------------------
Treatment: arm (3 groups)
PS variable: ps0
Reference group: 0
----------------------------------------------------------------------
-----------------------------------------------------------
Propensity Score Distribution
-----------------------------------------------------------
Placebo Low dose High dose
-----------------------------------------------------------
N 154 321 725
Mean 0.1515 0.2738 0.6163
SD 0.0608 0.0409 0.0831
Min 0.0422 0.1758 0.3046
Max 0.3586 0.4003 0.8513
-----------------------------------------------------------
-------------------------------------------------------
Common Support Region
-------------------------------------------------------
Lower bound: 0.3046
Upper bound: 0.3586
Outside support: 1129 (94.08%)
Placebo outside: 152
Low dose outside: 253
High dose outside: 724
-------------------------------------------------------
Warning: >10% of observations outside common support region.
Overlap: WARNING (94.1% outside support)
Consider: psdash support, threshold(0.05)
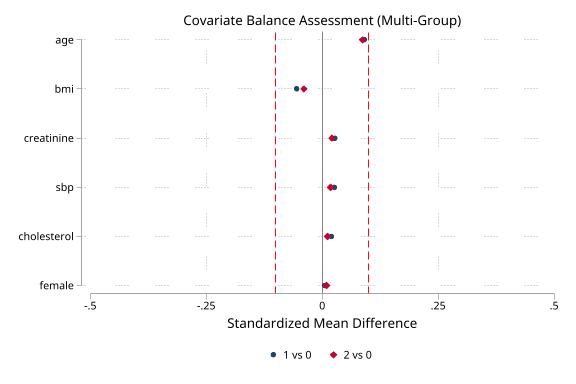
Multi-group balance (click to expand)
. noisily psdash balance arm, psvars(ps0 ps1 ps2)
> covariates(age female bmi sbp cholesterol creatinine) wvar(gipw)
---------------------------------------------------------------------------
Covariate Balance Assessment (Multi-Group)
---------------------------------------------------------------------------
Treatment: arm (3 groups, ref = 0)
Estimand: ATE
N (Group Placebo): 154
N (Group Low dose): 321
N (Group High dose): 725
Weights: gipw
Threshold: 0.100
---------------------------------------------------------------------------
-----------------------------------------------------------------------------------------------------
Covariate | SMD 1v0 VR SMD 2v0 VR Adj 1v0 VR Adj 2v0 VR Status
-----------------------------------------------------------------------------------------------------
age | 0.228 1.35 0.483 1.24 0.091 1.45 0.087 1.34 Balanced
female |-0.013 1.00-0.018 0.99 0.006 1.00 0.009 1.00 Balanced
bmi |-0.026 0.80 0.124 0.93-0.055 0.78-0.039 0.90 Balanced
sbp | 0.126 1.11 0.077 1.08 0.026 1.17 0.018 1.16 Balanced
cholesterol |-0.033 0.91-0.099 0.97 0.020 0.95 0.012 1.03 Balanced
creatinine |-0.231 0.87-0.269 0.85 0.028 0.89 0.021 0.85 Balanced
-----------------------------------------------------------------------------------------------------
Maximum |SMD| (raw): 0.483
Maximum |SMD| (adjusted): 0.091
Covariates > SMD threshold: 0 of 6
-----------------------------------------------------------------------------------------------------
Balance: Adequate (max |SMD| = 0.091)
Multi-group weight diagnostics (click to expand)
. noisily psdash weights arm, psvars(ps0 ps1 ps2) wvar(gipw)
----------------------------------------------------------------------
IPTW Weight Diagnostics (Multi-Group)
----------------------------------------------------------------------
Weight variable: gipw
Treatment: arm (3 groups, ref = 0)
Observations: 1,200
----------------------------------------------------------------------
-------------------------------------------------------------------------------------
Weight Distribution Summary
-------------------------------------------------------------------------------------
Overall Placebo Low dose High dose
-------------------------------------------------------------------------------------
N 1,200 154 321 725
Mean 2.989 7.708 3.737 1.655
SD 2.351 3.212 0.577 0.251
Min 1.175 2.789 2.498 1.175
Max 23.690 23.690 5.688 3.283
-------------------------------------------------------------------------------------
-------------------------------------------------------------------------------------
Effective Sample Size (ESS)
-------------------------------------------------------------------------------------
Overall Placebo Low dose High dose
-------------------------------------------------------------------------------------
ESS 741.5 131.3 313.6 708.8
ESS % of N 61.8% 85.3% 97.7% 97.8%
-------------------------------------------------------------------------------------
--------------------------------------------------
Extreme Weight Detection
--------------------------------------------------
Coefficient of Variation: 0.787
Weights > 10: 30 ( 2.50%)
Weights > 20: 1
--------------------------------------------------
Warning: 30 extreme weights detected (>10).
Warning: Maximum weight exceeds 20. Consider truncation.
Weights: Acceptable (ESS = 61.8% of N)
Multi-group common support (click to expand)
. noisily psdash support arm, psvars(ps0 ps1 ps2) threshold(0.1) nograph
----------------------------------------------------------------------
Common Support Assessment
----------------------------------------------------------------------
Treatment: arm (3 groups)
PS variable: ps0
Reference group: 0
Observations: 1,200
----------------------------------------------------------------------
-----------------------------------------------------------
Propensity Score Range
-----------------------------------------------------------
Placebo Low dose High dose
-----------------------------------------------------------
N 154 321 725
Min PS 0.0422 0.1758 0.3046
Max PS 0.3586 0.4003 0.8513
-----------------------------------------------------------
-------------------------------------------------------
Common Support Region
-------------------------------------------------------
Lower bound: 0.3046
Upper bound: 0.3586
Outside support: 1129 (94.08%)
Placebo outside: 152
Low dose outside: 253
High dose outside: 724
-------------------------------------------------------
-------------------------------------------------------
Manual Threshold Trimming
-------------------------------------------------------
Threshold: 0.1000
Trim region: [0.100, 0.900]
Observations trimmed: 30 ( 2.50%)
Remaining sample: 1170
-------------------------------------------------------
Warning: >10% of observations outside common support.
Support: Trimmed ( 2.5% excluded)
PNG demo images in demo/ are tracked in this repository.
Version History
- v1.0.0 (29 Apr 2026): Initial release with five subcommands